Installation
Please install from our r-universe:
install.packages(
"leapfrog",
repos = c("https://hivtools.r-universe.dev", "https://cloud.r-project.org"))You can install the latest release from GitHub with:
# install.packages("remotes")
remotes::install_github("hivtools/leapfrog", subdir = "leapfrogr", ref = "r-release")To install a development version on a branch checkout the repo to the appropriate branch and take the following steps:
-
(One time only) setup the
LEAPFROG_INCLUDEenv var using -
Generate the C++
-
Install the package
Simulation model
The “leapfrog” R package is an R interface around the “leapfrog” simulation model. The model is written in C++ and has interfaces in R, Python and C. Note the R package is in a subdirectory leapfrogr but the installed package is called leapfrog.
The simulation model is implemented in a header-only C++ library located in leapfrog-core/include/. The headers are bundled into inst/include in the installed package which allows the C++ code to be imported in other R packages via specifying LinkingTo: leapfrog in the DESCRIPTION file.
[!IMPORTANT] We use C++20 for this package. Please make sure you have a compiler that is compatible.
The simulation model is callable in R via a wrapper function run_model() created with Rcpp.
You can control how the simulation model is run by setting the configuration argument, see
leapfrog::list_model_configurations()for the list of available models. At the time of writing these are
-
DemographicProjection- runs on the demographic projections -
HivFullAgeStratification- runs the demographic projection and HIV adult projection with single-year age groups -
HivCoarseAgeStratification- runs the demographic projection and HIV adult projection with 5-year age groups -
ChildModel- runs the demographic projection, single-year age stratified HIV adult projection and HIV child projection
Example
The file inst/pjnz/bwa_aim-adult-art-no-special-elig_v6.13_2022-04-18.PJNZ contains an example Spectrum file constructed from default country data for Botswana with Spectrum (April 2022).
Prepare model inputs.
library(leapfrog)
pjnz <- system.file("pjnz/bwa_aim-adult-art-no-special-elig_v6.13_2022-04-18.PJNZ",
package = "leapfrog", mustWork = TRUE)
parameters <- process_pjnz(pjnz)
#> ℹ Some tags were not found in file:
#> • Tag not found in DP for `adult_non_aids_excess_mort`, returning `NULL`
#> • Tag not found in DP for `incidence_input`, returning `NULL`
#> • Tag not found in DP for `adult_art_adj_factor`, returning `NULL`
#> • Tag not found in DP for `adult_art_adj_factor_flag`, returning `NULL`
#> • Tag not found in DP for `adult_pats_alloc_to_from_other_region`, returning
#> `NULL`
#> • Tag not found in DP for `nosocomial_infections`, returning `NULL`
#> This message is displayed once per session.Simulate adult ‘full’ age group (single-year age) model from 1970 to 2030 with 10 HIV time steps per year.
lsimF <- run_model(parameters, "HivFullAgeStratification", 1970:2030)You can also simulate a model with ‘coarse’ age group (5-year age groups). You need to first prepare the coarse age group parameters
params_coarse <- process_pjnz(pjnz, use_coarse_age_groups = TRUE)
lsimC <- run_model(params_coarse, "HivCoarseAgeStratification", 1970:2030)Compare the HIV prevalence age 15-49 years and AIDS deaths 50+ years. Deaths 50+ years are to show some noticeable divergence between the "full" and "coarse" age group simulations.
prevF <- colSums(lsimF$p_hivpop[16:50,,],,2) / colSums(lsimF$p_totpop[16:50,,],,2)
prevC <- colSums(lsimC$p_hivpop[16:50,,],,2) / colSums(lsimC$p_totpop[16:50,,],,2)
deathsF <- colSums(lsimF$p_hiv_deaths[51:81,,],,2)
deathsC <- colSums(lsimC$p_hiv_deaths[51:81,,],,2)
plot(1970:2030, prevF, type = "l", main = "Prevalence 15-49")
lines(1970:2030, prevC, col = 2)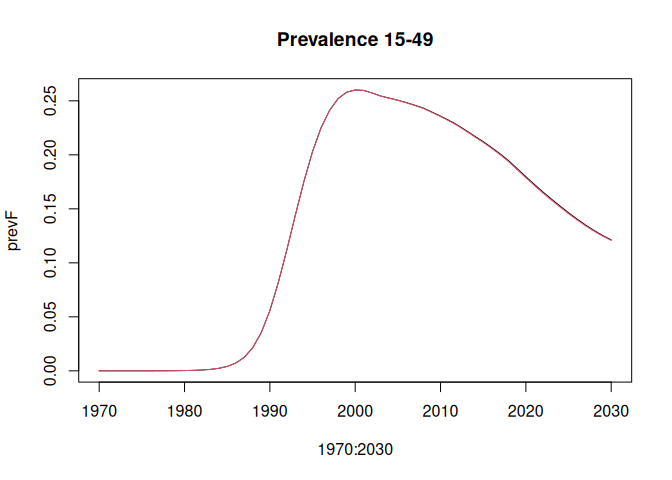
plot(1970:2030, deathsF, type = "l", main = "AIDS Deaths 50+ years")
lines(1970:2030, deathsC, col = 2)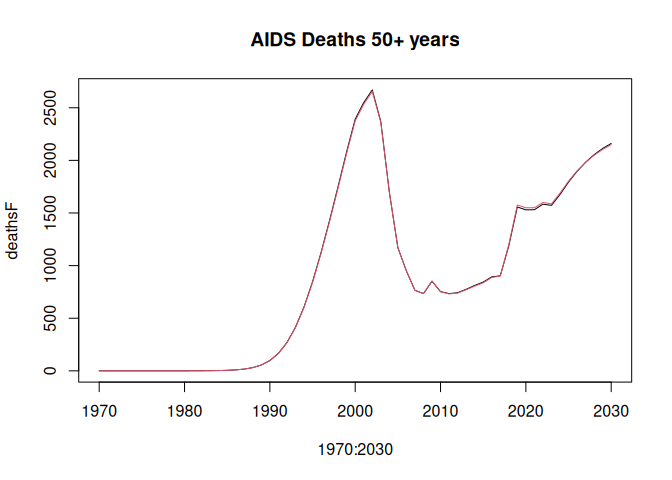
lint
Lint R code with lintr
lintr::lint_package()Lint C++ code with cpplint
cpplint leapfrog-core/include/*Code design
R functions
The file src/leapfrog.cpp contains R wrapper functions for the model simulation via Rcpp.
Development notes
Package loading
The C++ header library lives in leapfrog-core/include/ (outside the R package), with generated headers produced by a codegen step. How this is handled depends on how you are building/installing the package:
| Method |
configure runs in |
Headers used | Manual steps | Notes |
|---|---|---|---|---|
devtools::load_all() |
leapfrogr/ (source) |
LEAPFROG_INCLUDE or relative path to ../leapfrog-core/include
|
Run codegen ../scripts/generate manually on first run or when schemas change. |
When headers in leapfrog-core/include/ change, C++ recompiles automatically via Config/build/extra-sources. |
devtools::install() (default) |
Temp dir (tarball) |
LEAPFROG_INCLUDE env var |
LEAPFROG_INCLUDE must be set and run codegen manually |
configure runs from temp dir so ../leapfrog-core is absent. |
R CMD INSTALL leapfrogr/ |
leapfrogr/ (source) |
../leapfrog-core/include |
Nothing (in monorepo) | Will need LEAPFROG_INCLUDE env var if using R CMD build |
remotes::install_github(..., ref = "r-release") |
Temp dir (subdir only) |
inst/include (bundled) |
Nothing | Self-contained. r-release branch has headers bundled. Other branches will fail — use build=FALSE from a clone instead. |
| R-universe | Build server |
inst/include (bundled) |
Nothing | Preferred for end users. r-release branch is used per packages.json. Updated on every merge to main. |
To set up LEAPFROG_INCLUDE for local development, run the helper script once:
Note that LEAPFROG_INCLUDE is not needed for remotes::install_github(..., ref = "r-release") or r-universe installs — those use the r-release branch which has the C++ headers bundled into inst/include/. The r-release branch is updated automatically on every merge to main.
For CRAN submission (when we add it), build the tarball from the r-release branch — the headers are already present so it will pass R CMD check.
Testing
There is some pre-prepared test data available to make tests run faster. This is generated and saved by ../scripts/create_test_data.R as hdf5 format files.
This data is used in leapfrog-py for pythont tests and cpp_interface for testing standalone C++ model.